Software to protect the world's most endangered species

Sheep grazing near Mount Atlas, in northern Morocco. © iStock
By combining genetic and environmental databases, researchers at EPFL are seeking to help biologists identify more accurately the animal and plant species most exposed to climate change, in order to develop appropriate conservation methods.
Northern Morocco is home to a type of sheep that has a specific gene, developed over thousands of years of evolution. The gene makes the sheep secrete a type of wax from its coat, protecting it from the heavy precipitation in the High Atlas region where it lives. By preventing the sheep’s coat from rotting, the gene protects it against potentially fatal skin diseases.
At EPFL, researchers in the Laboratory of Geographic Information Systems (LASIG) have developed software to assist evolutionary biologists. By providing them with simultaneous and direct access to bioinformatics databases featuring automatic genome annotations and to climate databases, it saves them weeks of work. Climate databases contain information about precipitation, wind, sunlight and cloud cover in a given region. The software, named R.Samada, assesses the relationship between the genetic and environmental information and generates graphs and maps that allow researchers to visualize the data rapidly. The software was unveiled in a paper published in Molecular Ecology Resources.
From genes to maps
When they generate a map of Morocco showing the location of sheep featuring the genetic marker that protects them from rain, biologists will see that a certain variant of this gene is only present in sheep living in the mountainous areas of the Atlas region. In the deserts of southern Morocco, sheep lack this genetic marker. A climate scenario for 2070 in which temperatures rise by 3.7°C – based on figures from the Intergovernmental Panel on Climate Change (IPCC) – shows that precipitation will decrease in this region, and so it is very likely that many sheep will need another genetic variant to adapt to the coming drought.
“The software identifies the genes involved in the process through which a species evolves to adapt to weather conditions”, explains Stéphane Joost, corresponding author of the paper and thesis supervisor of Solange Duruz, first author. “With this software, we wanted to give evolutionary and conservation biologists a single, simplified point of access to all of the processing and analysis of genetic and geo-environmental information commonly used to assess the status of species, particularly those whose survival is threatened by climate change.”
Making the right decisions
What will happen to the Moroccan sheep as the climate changes? What if there is less rainfall where they live? As the planet becomes warmer and as biodiversity decreases, evolutionary biologists are pursuing two parallel aims: they want to cryopreserve the DNA of species threatened with extinction and identify the environments in which a species is most likely to survive in a given climate scenario. The software developed by EPFL will help them make the right decisions by showing how well genetic variants fit with certain eco-climatic zones.
Comparing results
R.Sambada is the latest version of a system that has been under development at EPFL for more than ten years. It is one of five similar software packages available in the market showing varying levels of sophistication. R.Samada is currently the fastest of them. “We are encouraging researchers to compare our software’s results with those of other systems, in order to make their analysis and methods more robust,” adds Joost.
Through research currently being done for a PhD thesis, EPFL intends to develop its software further, with the particular aim of generating conservation zones based on various climate scenarios involving temperature increases of between 1.5°C and 4°C between now and 2100. “Once the software has this feature, we will be able to more accurately pinpoint the zones at risk and those that could provide refuge,” concludes Joost. Work is also planned to adapt the system to marine zones, with a view to preserving coral for example, and to improve map resolution.
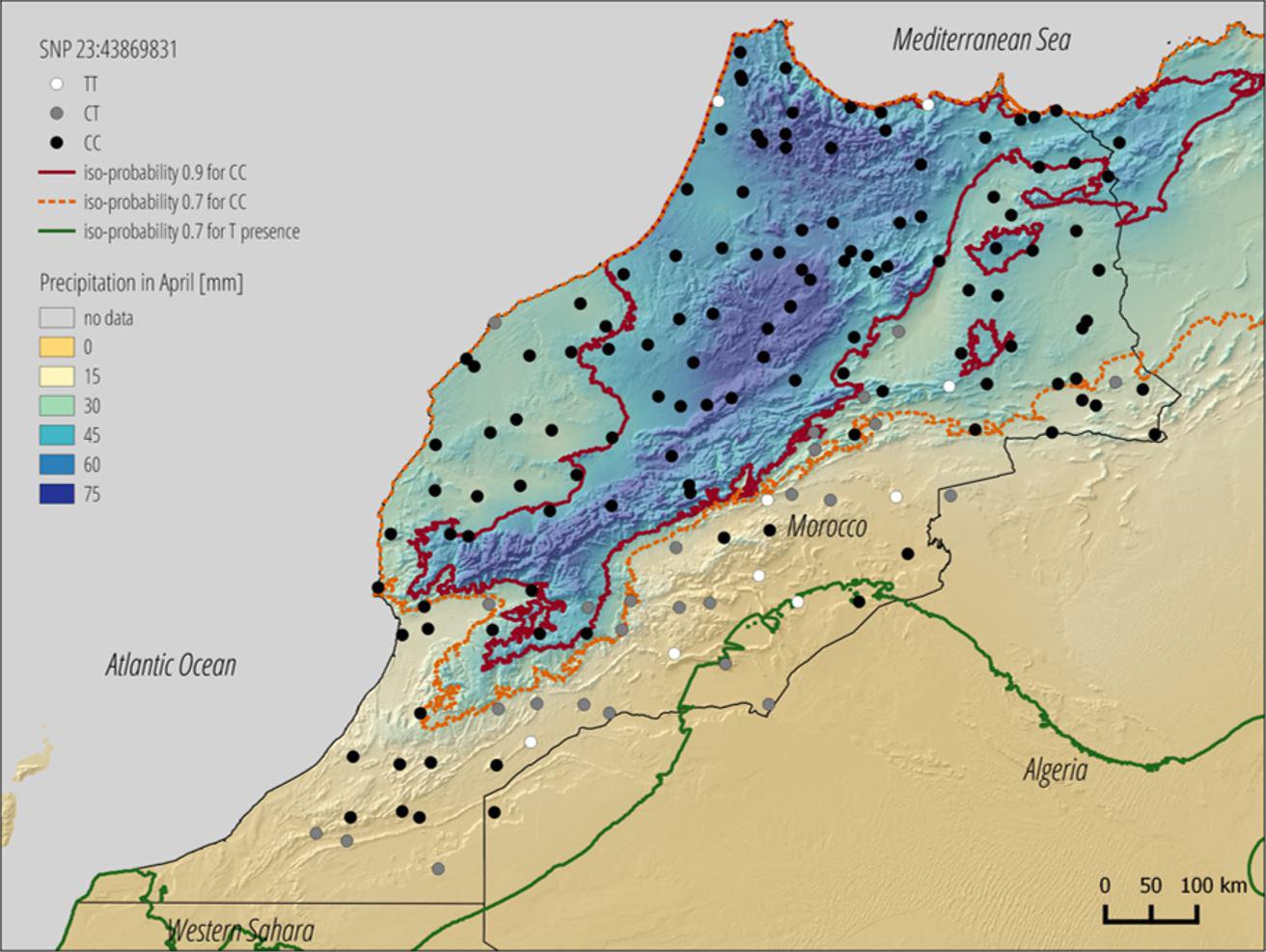 © LASIG / EPFL
© LASIG / EPFL
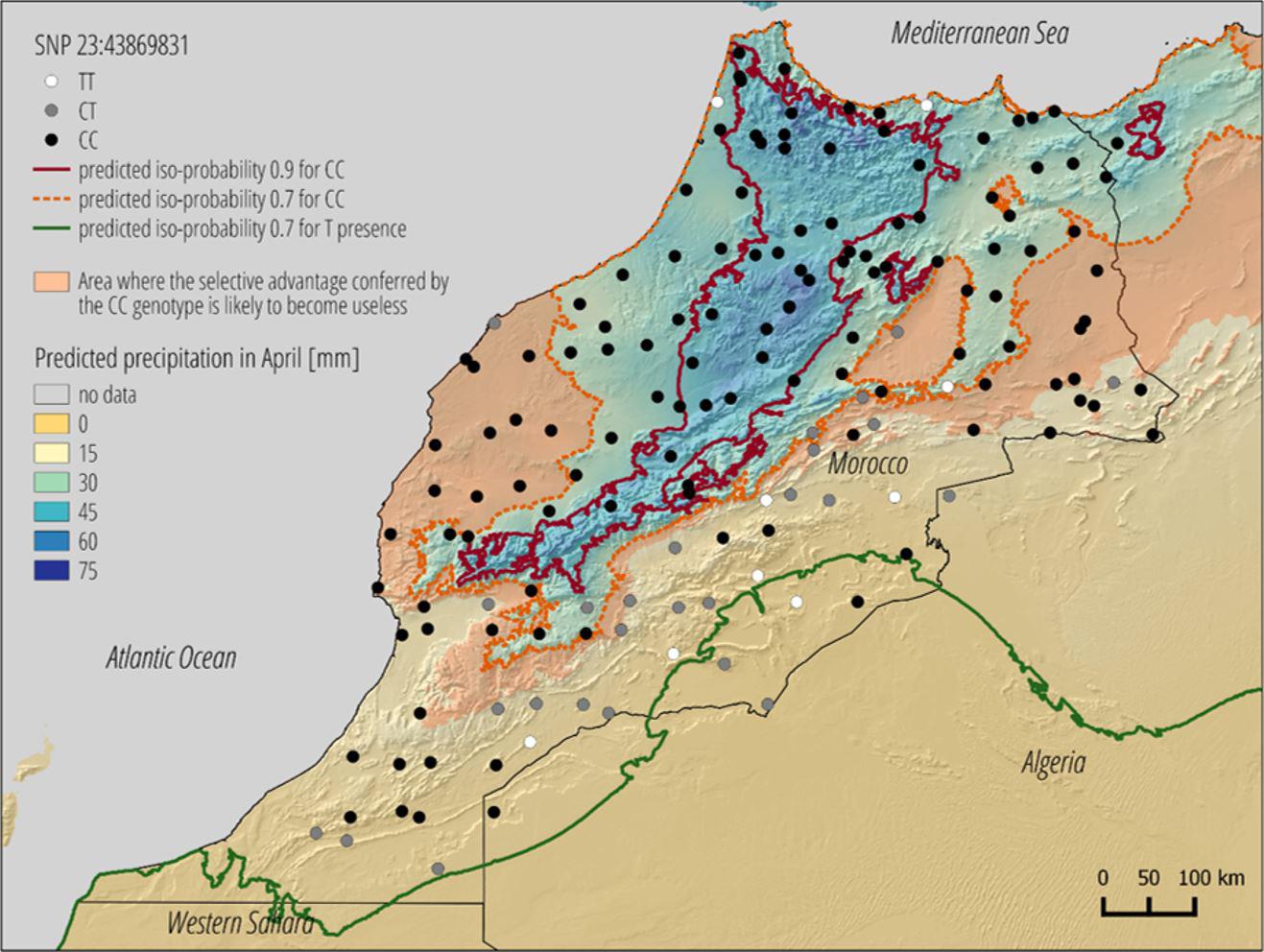 © LASIG / EPFL
© LASIG / EPFL

